Note
Go to the end to download the full example code
Rb₂SO₄, ⁸⁷Rb (I=3/2) SAS¶
⁸⁷Rb (I=3/2) Switched-angle spinning (SAS) simulation.
The following is an example of Switched-Angle Spinning (SAS) simulation of \(\text{Rb}_2\text{SO}_4\), which has two distinct rubidium sites. The NMR tensor parameters for these sites are taken from Shore et al. [1].
import numpy as np
import matplotlib.pyplot as plt
from mrsimulator import Simulator, SpinSystem, Site
from mrsimulator.method import Method, SpectralDimension, SpectralEvent
from mrsimulator import signal_processor as sp
from mrsimulator.spin_system.tensors import SymmetricTensor
Generate the site and spin system objects.
sites = [
Site(
isotope="87Rb",
isotropic_chemical_shift=16, # in ppm
quadrupolar=SymmetricTensor(Cq=5.3e6, eta=0.1), # Cq in Hz
),
Site(
isotope="87Rb",
isotropic_chemical_shift=40, # in ppm
quadrupolar=SymmetricTensor(Cq=2.6e6, eta=1.0), # Cq in Hz
),
]
spin_systems = [SpinSystem(sites=[s]) for s in sites]
Use the generic Method class to simulate a 2D SAS spectrum by customizing the method parameters, as shown below.
sas = Method(
name="Switched Angle Spinning",
channels=["87Rb"],
magnetic_flux_density=9.4, # in T
rotor_frequency=np.inf,
spectral_dimensions=[
SpectralDimension(
count=256,
spectral_width=3.5e4, # in Hz
reference_offset=1e3, # in Hz
label="90 dimension",
events=[
SpectralEvent(
rotor_angle=90 * np.pi / 180, # in radians
transition_queries=[{"ch1": {"P": [-1], "D": [0]}}],
)
],
),
SpectralDimension(
count=256,
spectral_width=22e3, # in Hz
reference_offset=-4e3, # in Hz
label="MAS dimension",
events=[
SpectralEvent(
rotor_angle=54.74 * np.pi / 180, # in radians
transition_queries=[{"ch1": {"P": [-1], "D": [0]}}],
)
],
),
],
)
# A graphical representation of the method object.
plt.figure(figsize=(5, 2.5))
sas.plot()
plt.show()
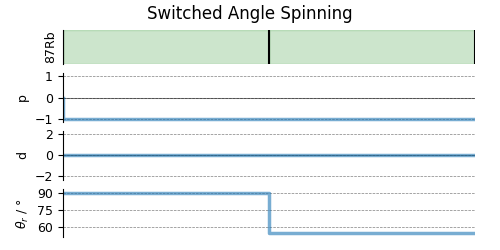
Create the Simulator object, add the method and spin system objects, and run the simulation.
The plot of the simulation.
dataset = sim.methods[0].simulation
plt.figure(figsize=(4.25, 3.0))
ax = plt.subplot(projection="csdm")
cb = ax.imshow(dataset.real / dataset.real.max(), aspect="auto", cmap="gist_ncar_r")
plt.colorbar(cb)
ax.invert_xaxis()
plt.tight_layout()
plt.show()
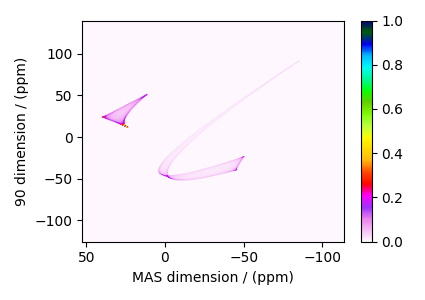
Add post-simulation signal processing.
processor = sp.SignalProcessor(
operations=[
# Gaussian convolution along both dimensions.
sp.IFFT(dim_index=(0, 1)),
sp.apodization.Gaussian(FWHM="0.4 kHz", dim_index=0),
sp.apodization.Gaussian(FWHM="0.4 kHz", dim_index=1),
sp.FFT(dim_index=(0, 1)),
]
)
processed_dataset = processor.apply_operations(dataset=dataset)
processed_dataset /= processed_dataset.max()
The plot of the simulation after signal processing.
plt.figure(figsize=(4.25, 3.0))
ax = plt.subplot(projection="csdm")
cb = ax.imshow(processed_dataset.real, cmap="gist_ncar_r", aspect="auto")
plt.colorbar(cb)
ax.invert_xaxis()
plt.tight_layout()
plt.show()
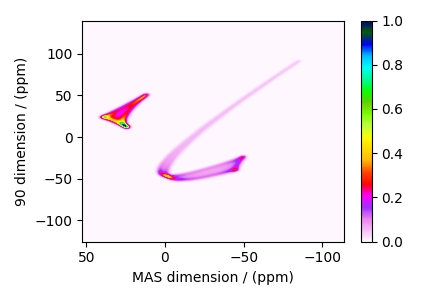
Total running time of the script: (0 minutes 1.100 seconds)
