Methods Library¶
For convenience, mrsimulator offers the following pre-built methods -
An example of the syntax that all library methods follows is shown below.
from mrsimulator.method import SpectralDimension
from mrsimulator.method.lib import BlochDecaySpectrum
lib_method = BlochDecaySpectrum(
channels=["29Si"], # list of isotopes
magnetic_flux_density=4.7, # T
rotor_angle=57.735 * 3.1415 / 180, # rad
rotor_frequency=10000, # Hz
spectral_dimensions=[
SpectralDimension(count=512, spectral_width=5e4, reference_offset=10),
],
)
where BlochDecaySpectrum can be replaced with another library method class. Each method has the
channels attribute, which is a list of isotopes probed by the method as well as the
magnetic_flux_density, rotor_angle, and rotor_frequency attributes which define the
global experiment parameters.
See Table 6 for more details.
The method object also has the spectral_dimensions attribute, which contains a list of
SpectralDimension objects defining the spectral grid. A 2D method will have two spectral
dimensions in this list, whereas a 1D method will only have one. See
Table 7 for the attributes of a SpectralDimension object.
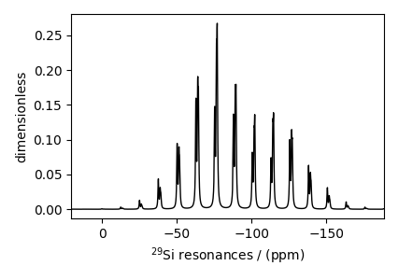
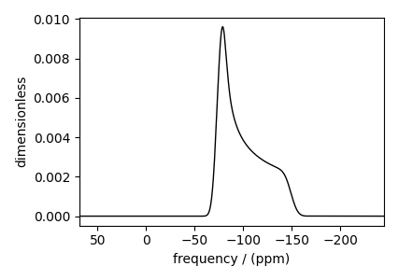
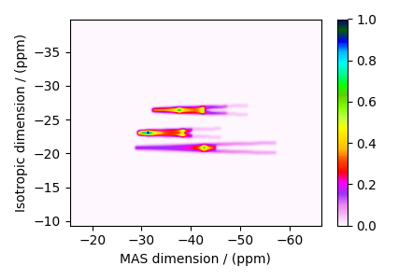
Bloch Decay Spectrum¶
The BlochDecaySpectrum class simulates the
Bloch decay spectrum.
from mrsimulator.method.lib import BlochDecaySpectrum
from mrsimulator.method import SpectralDimension
method = BlochDecaySpectrum(
channels=["1H"],
rotor_frequency=12500, # in Hz
rotor_angle=54.735 * 3.14159 / 180, # in rad
magnetic_flux_density=9.4, # in tesla
spectral_dimensions=[
SpectralDimension(
count=1024,
spectral_width=25e3, # in Hz
reference_offset=-4e3, # in Hz
)
],
)
Examples
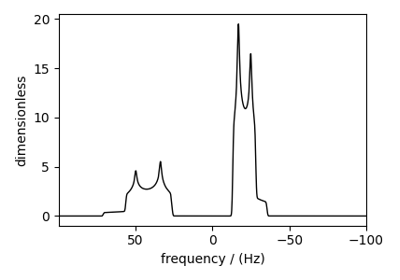
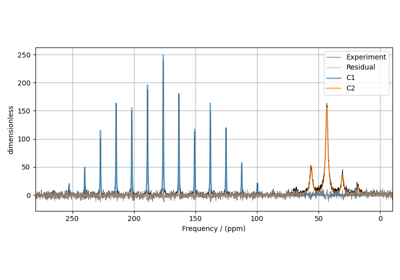
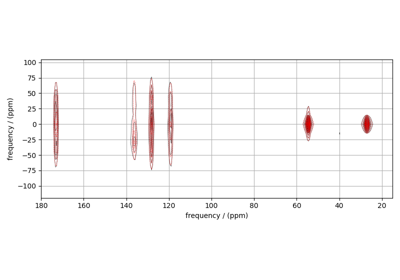
Influence of 14N on 13C NMR MAS spectra of glycine
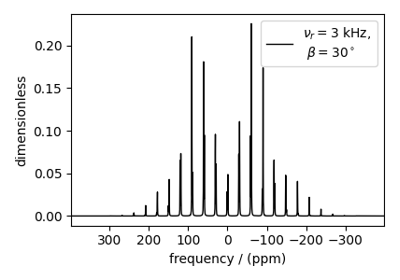
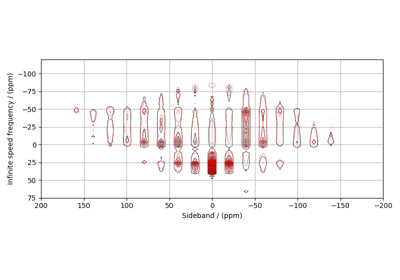
Coupled spin-1/2 (CSA + heteronuclear dipolar + J-couplings)
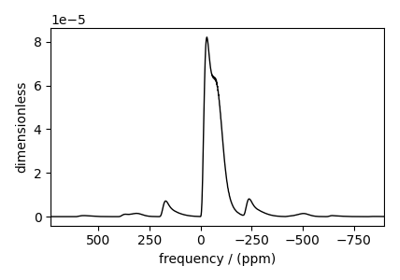
Extended Czjzek distribution (Shielding and Quadrupolar)
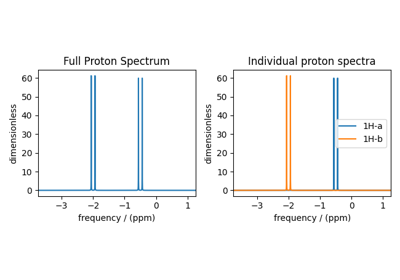
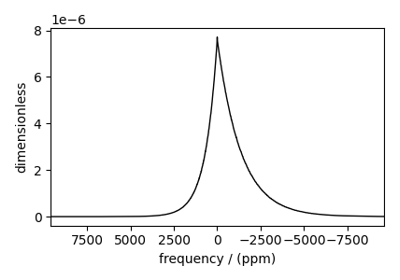
Bloch Decay Central Transition¶
The BlochDecayCTSpectrum class simulates the
Bloch decay central transition selective spectrum.
from mrsimulator.method.lib import BlochDecayCTSpectrum
from mrsimulator.method import SpectralDimension
method = BlochDecayCTSpectrum(
channels=["87Rb"],
rotor_frequency=12500, # in Hz
rotor_angle=54.735 * 3.14159 / 180, # in rad
magnetic_flux_density=9.4, # in tesla
spectral_dimensions=[
SpectralDimension(
count=1024,
spectral_width=25e3, # in Hz
reference_offset=-4e3, # in Hz
)
],
)
Examples

Extended Czjzek distribution (Shielding and Quadrupolar)
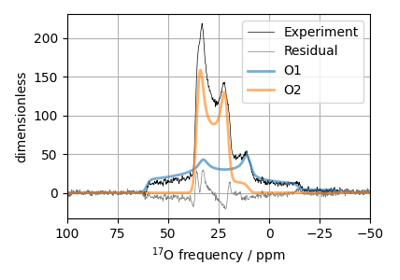
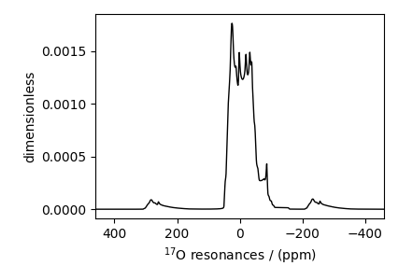
¹⁷O MAS NMR of crystalline Na₂SiO₃ (2nd order quad)
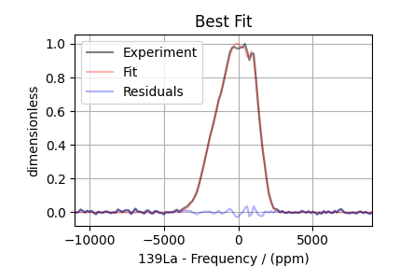
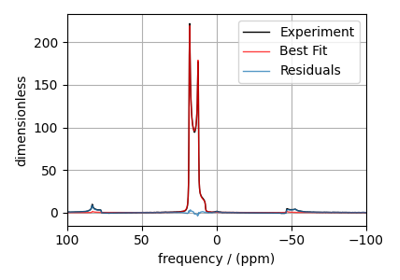
Extended Czjzek fitting of ¹³⁹La MAS NMR of La₀.₂Y₁.₈Si₂2O₇
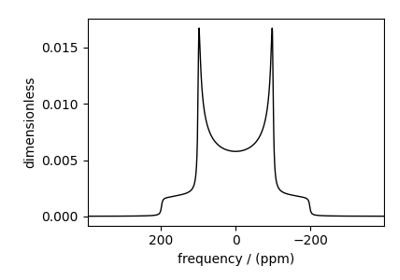
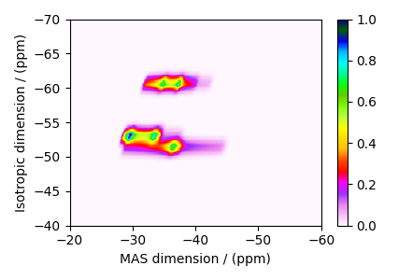
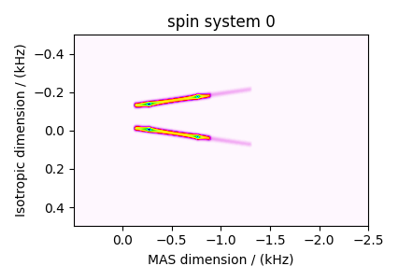
Multi-Quantum VAS¶
The ThreeQ_VAS, FiveQ_VAS, and
SevenQ_VAS classes all simulate a multiple quantum VAS
spectrum. The spinning speed for all three methods is fixed at infinite speed. The spectrum
is also sheared such that the correlated dimensions are the isotropic dimension and the VAS dimension.
from mrsimulator.method.lib import ThreeQ_VAS
from mrsimulator.method import SpectralDimension
method = ThreeQ_VAS(
channels=["87Rb"],
magnetic_flux_density=7, # in T
spectral_dimensions=[
SpectralDimension(
count=128,
spectral_width=3e3, # in Hz
reference_offset=-2e3, # in Hz
label="Isotropic dimension",
),
SpectralDimension(
count=512,
spectral_width=1e4, # in Hz
reference_offset=-5e3, # in Hz
label="MAS dimension",
),
],
)
The other methods, employing five and seven quantum transitions, can be imported as follows:
from mrsimulator.method.lib import FiveQ_VAS
from mrsimulator.method.lib import SevenQ_VAS
Examples
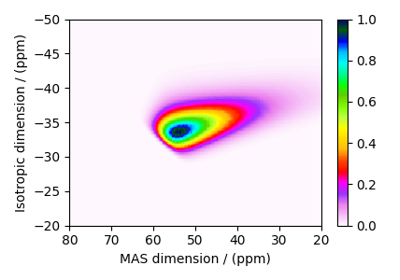
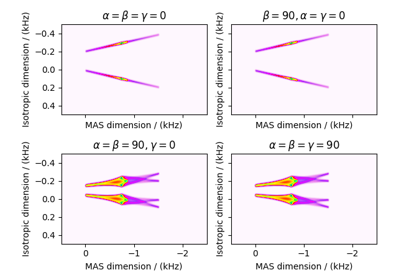
Satellite-Transition VAS¶
The ST1_VAS and ST2_VAS classes
simulate a sheared and scaled satellite and central transition correlation spectrum. The spinning
speed for these methods is fixed at infinite speed.
from mrsimulator.method.lib import ST1_VAS
from mrsimulator.method import SpectralDimension
method = ST1_VAS(
channels=["87Rb"],
magnetic_flux_density=9.4, # in T
spectral_dimensions=[
SpectralDimension(
count=128,
spectral_width=1e3, # in Hz
reference_offset=-5e3, # in Hz
label="Isotropic dimension",
),
SpectralDimension(
count=256,
spectral_width=1e4, # in Hz
reference_offset=-3e3, # in Hz
label="MAS dimension",
),
],
)
Examples
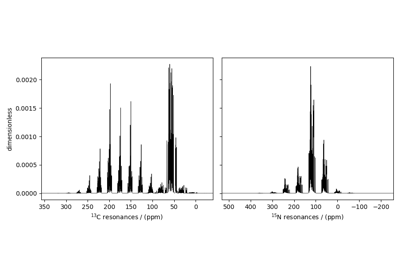
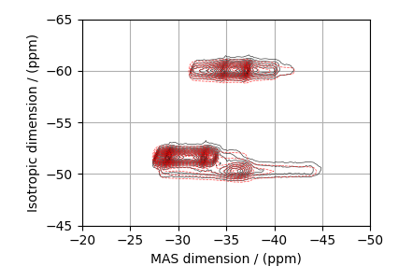
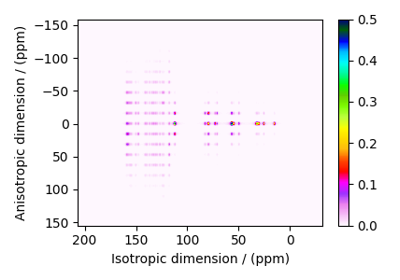
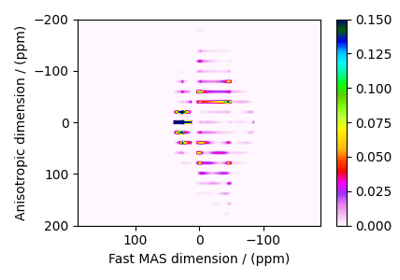
SSB2D¶
The SSB2D class simulates a sheared 2D finite
to infinite speed MAS correlation spectrum. The spinning speed for the second spectral
dimension is fixed at infinite spinning speed
from mrsimulator.method.lib import SSB2D
from mrsimulator.method import SpectralDimension
method = SSB2D(
channels=["13C"],
magnetic_flux_density=7, # in T
rotor_frequency=1500, # in Hz
spectral_dimensions=[
SpectralDimension(
count=16,
spectral_width=16 * 1500, # in Hz (= count * rotor_frequency)
reference_offset=-5e3, # in Hz
label="Sideband dimension",
),
SpectralDimension(
count=512,
spectral_width=1e4, # in Hz
reference_offset=-4e3, # in Hz
label="Isotropic dimension",
),
],
)
Examples
Attribute Summaries¶
Attribute Name |
Type |
Description |
|---|---|---|
channels |
|
A required list of isotopes given as strings over which the given method applies.
For example, |
magnetic_flux_density |
|
An optional float describing the macroscopic magnetic flux density of the applied
external magnetic field in tesla. For example, |
rotor_frequency |
|
An optional float describing the sample rotation frequency in Hz. For example, |
rotor_angle |
|
An optional float describing the angle between the sample rotation axis and the external
magnetic field in radians. The default value is the magic angle,
|
spectral_dimensions |
|
A list of SpectralDimension objects describing the spectral dimensions for the method. |
simulation |
CSDM object |
A CSDM object representing the spectrum simulated by the method. By default, the value is
|
experiment |
CSDM object |
An optional CSDM object holding an experimental measurement of the method. The default
value is |
Attribute Name |
Type |
Description |
|---|---|---|
count |
|
An optional integer representing the number of points, \(N\), along the spectroscopic
dimension. For example, |
spectral_width |
|
An optional float representing the width, \(\Delta x\), of the spectroscopic dimension
in Hz. For example, |
reference_offset |
|
An optional float representing the reference offset, \(x_0\), of the spectroscopic
dimension in Hz. For example, |
origin_offset |
|
An optional float representing the origin offset, or Larmor frequency, along the
spectroscopic dimension in units of Hz. The default value is |