Note
Go to the end to download the full example code
NiCl₂.2D₂O, ²H (I=1) Shifting-d echo¶
²H (I=1) 2D NMR CSA-Quad 1st order correlation spectrum.
The following is an example of fitting static shifting-d echo NMR correlation spectrum of \(\text{NiCl}_2\cdot 2\text{D}_2\text{O}\) crystalline solid. The spectrum used here is from Walder et al. [1].
import numpy as np
import csdmpy as cp
import matplotlib.pyplot as plt
from lmfit import Minimizer
from mrsimulator import Simulator, Site, SpinSystem
from mrsimulator import signal_processor as sp
from mrsimulator.utils import spectral_fitting as sf
from mrsimulator.utils import get_spectral_dimensions
from mrsimulator.spin_system.tensors import SymmetricTensor
from mrsimulator.method import Method, SpectralDimension, SpectralEvent, MixingEvent
Import the dataset¶
filename = "https://ssnmr.org/sites/default/files/mrsimulator/NiCl2.2D2O.csdf"
experiment = cp.load(filename)
# For spectral fitting, we only focus on the real part of the complex dataset
experiment = experiment.real
# Convert the coordinates along each dimension from Hz to ppm.
_ = [item.to("ppm", "nmr_frequency_ratio") for item in experiment.dimensions]
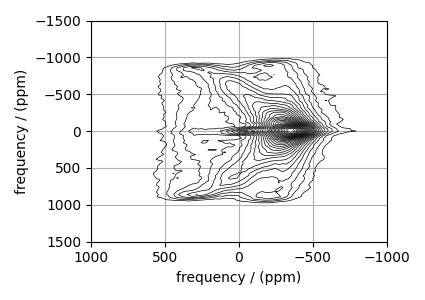
# plot of the dataset.
max_amp = experiment.max()
levels = (np.arange(29) + 1) * max_amp / 30 # contours are drawn at these levels.
options = dict(levels=levels, linewidths=0.5) # plot options
plt.figure(figsize=(4.25, 3.0))
ax = plt.subplot(projection="csdm")
ax.contour(experiment, colors="k", **options)
ax.set_xlim(1000, -1000)
ax.set_ylim(1500, -1500)
plt.grid()
plt.tight_layout()
plt.show()
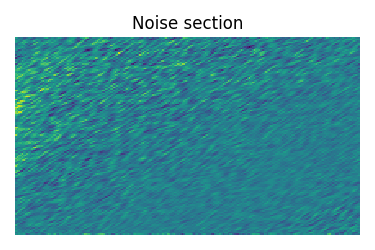
Estimate noise statistics from the dataset
coords = experiment.dimensions[0].coordinates
noise_region = np.where(coords > 700e-6)
noise_data = experiment[noise_region]
plt.figure(figsize=(3.75, 2.5))
ax = plt.subplot(projection="csdm")
ax.imshow(noise_data, aspect="auto", interpolation="none")
plt.title("Noise section")
plt.axis("off")
plt.tight_layout()
plt.show()
noise_mean, sigma = experiment[noise_region].mean(), experiment[noise_region].std()
noise_mean, sigma
(<Quantity 0.64582>, <Quantity 5.835975>)
Create a fitting model¶
Guess model
Create a guess list of spin systems.
site = Site(
isotope="2H",
isotropic_chemical_shift=-90, # in ppm
shielding_symmetric=SymmetricTensor(
zeta=-610, # in ppm
eta=0.15,
alpha=0.7, # in rads
beta=2.0, # in rads
gamma=3.0, # in rads
),
quadrupolar=SymmetricTensor(Cq=75.2e3, eta=0.9), # Cq in Hz
)
spin_systems = [SpinSystem(sites=[site])]
Method
Use the generic method, Method, to generate a shifting-d echo method. The
reported shifting-d 2D sequence is a correlation of the shielding frequencies to the
first-order quadrupolar frequencies. Here, we create a correlation method using the
freq_contrib attribute, which acts as a switch
for including the frequency contributions from interaction during the event.
In the following method, we assign the ["Quad1_2"] and
["Shielding1_0", "Shielding1_2"] as the value to the freq_contrib key. The
Quad1_2 is an enumeration for selecting the first-order second-rank quadrupolar
frequency contributions. Shielding1_0 and Shielding1_2 are enumerations for
the first-order shielding with zeroth and second-rank tensor contributions,
respectively. See FrequencyEnum for details.
# Get the spectral dimension parameters from the experiment.
spectral_dims = get_spectral_dimensions(experiment)
shifting_d = Method(
channels=["2H"],
magnetic_flux_density=9.395, # in T
rotor_frequency=0, # in Hz
rotor_angle=0, # in rads
spectral_dimensions=[
SpectralDimension(
**spectral_dims[0],
label="Quadrupolar frequency",
events=[
SpectralEvent(
transition_queries=[{"ch1": {"P": [-1]}}],
freq_contrib=["Quad1_2"],
),
MixingEvent(query="NoMixing"),
],
),
SpectralDimension(
**spectral_dims[1],
label="Paramagnetic shift",
events=[
SpectralEvent(
transition_queries=[{"ch1": {"P": [-1]}}],
freq_contrib=["Shielding1_0", "Shielding1_2"],
)
],
),
],
experiment=experiment, # also add the measurement to the method.
)
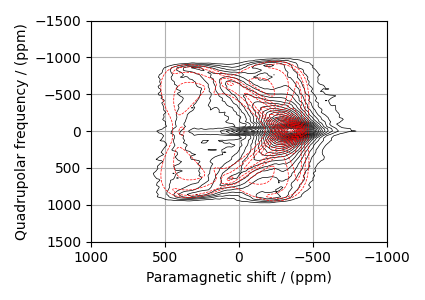
Guess Spectrum
# Simulation
# ----------
sim = Simulator(spin_systems=spin_systems, methods=[shifting_d])
sim.config.integration_volume = "hemisphere"
sim.run()
# Post Simulation Processing
# --------------------------
processor = sp.SignalProcessor(
operations=[
# Gaussian convolution along both dimensions.
sp.IFFT(dim_index=(0, 1)),
sp.apodization.Gaussian(FWHM="5 kHz", dim_index=0), # along dimension 0
sp.apodization.Gaussian(FWHM="5 kHz", dim_index=1), # along dimension 1
sp.FFT(dim_index=(0, 1)),
sp.Scale(factor=5e9),
]
)
processed_dataset = processor.apply_operations(dataset=sim.methods[0].simulation).real
# Plot of the guess Spectrum
# --------------------------
plt.figure(figsize=(4.25, 3.0))
ax = plt.subplot(projection="csdm")
ax.contour(experiment, colors="k", **options)
ax.contour(processed_dataset, colors="r", linestyles="--", **options)
ax.set_xlim(1000, -1000)
ax.set_ylim(1500, -1500)
plt.grid()
plt.tight_layout()
plt.show()
Least-squares minimization with LMFIT¶
Use the make_LMFIT_params() for a quick
setup of the fitting parameters.
params = sf.make_LMFIT_params(sim, processor)
print(params.pretty_print(columns=["value", "min", "max", "vary", "expr"]))
Name Value Min Max Vary Expr
SP_0_operation_1_Gaussian_FWHM 5 -inf inf True None
SP_0_operation_2_Gaussian_FWHM 5 -inf inf True None
SP_0_operation_4_Scale_factor 5e+09 -inf inf True None
sys_0_abundance 100 0 100 False 100
sys_0_site_0_isotropic_chemical_shift -90 -inf inf True None
sys_0_site_0_quadrupolar_Cq 7.52e+04 -inf inf True None
sys_0_site_0_quadrupolar_eta 0.9 0 1 True None
sys_0_site_0_shielding_symmetric_alpha 0.7 -inf inf True None
sys_0_site_0_shielding_symmetric_beta 2 -inf inf True None
sys_0_site_0_shielding_symmetric_eta 0.15 0 1 True None
sys_0_site_0_shielding_symmetric_gamma 3 -inf inf True None
sys_0_site_0_shielding_symmetric_zeta -610 -inf inf True None
None
Solve the minimizer using LMFIT
opt = sim.optimize() # Pre-compute transition pathways
minner = Minimizer(
sf.LMFIT_min_function,
params,
fcn_args=(sim, processor, sigma),
fcn_kws={"opt": opt},
)
result = minner.minimize()
result
The best fit solution¶
best_fit = sf.bestfit(sim, processor)[0].real
# Plot the spectrum
plt.figure(figsize=(4.25, 3.0))
ax = plt.subplot(projection="csdm")
ax.contour(experiment, colors="k", **options)
ax.contour(best_fit, colors="r", linestyles="--", **options)
ax.set_xlim(1000, -1000)
ax.set_ylim(1500, -1500)
plt.grid()
plt.tight_layout()
plt.show()
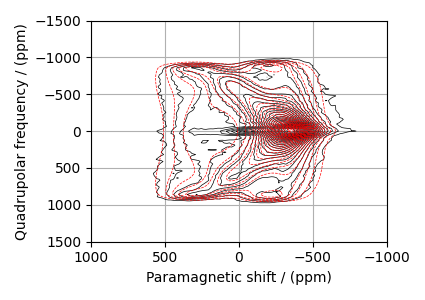
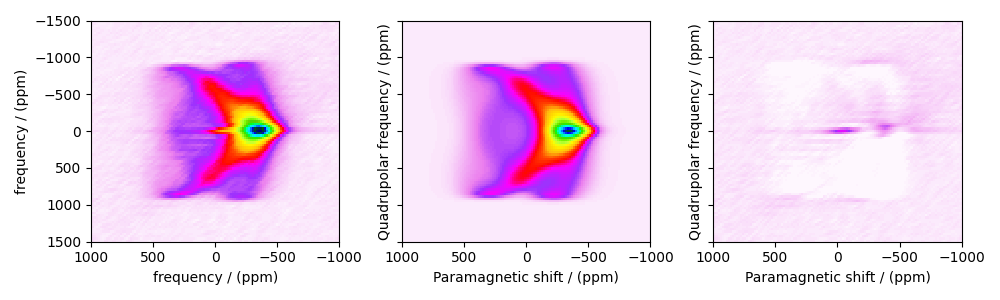
Image plots with residuals¶
residuals = sf.residuals(sim, processor)[0].real
fig, ax = plt.subplots(
1, 3, sharey=True, figsize=(10, 3.0), subplot_kw={"projection": "csdm"}
)
vmax, vmin = experiment.max(), experiment.min()
for i, dat in enumerate([experiment, best_fit, residuals]):
ax[i].imshow(
dat,
aspect="auto",
cmap="gist_ncar_r",
vmax=vmax,
vmin=vmin,
interpolation="none",
)
ax[i].set_xlim(1000, -1000)
ax[0].set_ylim(1500, -1500)
plt.tight_layout()
plt.show()
Total running time of the script: (0 minutes 11.851 seconds)
