Single Site System Generator¶
Custom Site and SpinSystem parameters can be passed to the
single_site_system_generator() method to create a list
of uncoupled spin systems. Each SpinSystem in the returned list holds a single
Site object since the backend simulation is more efficient for single site spin
systems. Import the method as below
from mrsimulator.utils.collection import single_site_system_generator
The arguments passed to the function, defined in Table 5,
can either be a scalar quantity (float or str, where applicable) or a
list/np.array of those quantities. All lists passed must
have the same length, otherwise an error will be thrown. For example,
single_site_system_generator(
isotope=["1H", "1H", "13C", "17O"],
isotropic_chemical_shift=[1.3, 3.7, 65.0],
)
Out:
Traceback (most recent call last):
...
ValueError: An array or list was either too short or too long. All arguments must be the
same size. If one attribute is a type list of length n, then all attributes with list types
must also be of length n, and all remaining attributes must be scalar (singular float, int,
or str).
The attributes of each returned spin system at a certain index correspond to the attribute passed at that index. For example,
single_site_system_generator(
isotope=["1H", "1H", "13C"],
isotropic_chemical_shift=[1.3, 3.7, 65.0],
)
returns a list of 3 spin systems. The first two spin systems represent proton sites with isotropic chemical shifts of 1.3 ppm and 3.7 ppm, respectively. The third spin system is a \(^{13}\text{C}\) site with a chemical shift of 65.0 ppm.
Broadcasting Length of List¶
Arguments passed as a single value will be broadcast to a list of that value with the same length as other lists passed. For example
single_site_system_generator(
isotope=["1H", "1H", "1H"],
isotropic_chemical_shift=2.0,
)
is equivalent to calling
single_site_system_generator(
isotope=["1H", "1H", "1H"],
isotropic_chemical_shift=[2.0, 2.0, 2.0],
)
Passing lists of Tensor Parameters¶
Tensor parameters for sites are passed as dictionaries where the keywords represent the tensor
attribute and the values are single values or a list/np.array of values. Again, these
lists must have the same length of all other lists passed. Single values will be broadcast to a
list of that value with the same length as other lists passed. For example
single_site_system_generator(
isotope="13C",
shielding_symmetric={
"zeta": [5, 10, 15, 20, 25],
"eta": 0.3,
},
)
returns a list of five \(^{13}\text{C}\) spin systems with different shielding_symmetric.zeta
values but the same shielding_symmetric.eta value.
If you need to intermix sites with and without tensor parameters, simply put None at the index
of the site without the tensor parameter.
single_site_system_generator(
isotope=["1H", "17O"],
quadrupolar={
"Cq": [None, 3.2e6],
"eta": [None, 0.5],
},
)
Examples using single_site_system_generator()¶
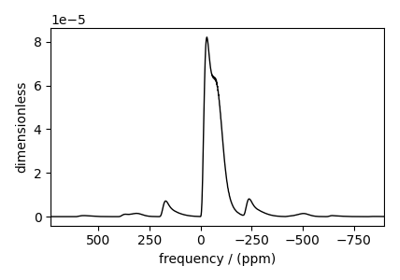
Extended Czjzek distribution (Shielding and Quadrupolar)
Name |
Type |
Description |
|---|---|---|
|
|
A required string or list of strings representing the label of the |
|
|
An optional number or list of numbers representing the |
|
|
An optional dictionary representing the |
|
|
An optional dictionary representing the |
|
|
An optional dictionary representing the |
|
|
An optional number or list of numbers representing the |
|
|
An optional string or list of strings representing the |
|
|
An optional string or list of strings representing the |
|
|
An optional string or list of strings representing the |